 
While OE-PCR provides efficient methods for introducing multiple and large changes, they involve multiple rounds of PCR and DNA purification, limiting the creation of multiple mutations simultaneously. Recently, several labs reported alternatives, such as overlap extension PCR (OE-PCR) 13, 14, 15, 16 and homologous recombination-based methods 17, 18, 19, 20, 21, 22, 23, 24, 25, for creating mutations in vitro or in vivo. Despite high efficiency, these approaches require primers containing the desired mutations in the template annealing regions, which limits the introduction of large changes required in some functional studies. These methods use partially overlapping primers to reduce the formation of primer dimers and hence improve PCR amplification efficiency. To circumvent these limitations, many modified versions of the QuikChange site-directed mutagenesis method have been developed 4, 10, 11, 12. In addition, the originally developed QuikChange method requires the altered nucleotides to be introduced in the middle of both primers, limiting the introduction of multiple mutations 4 as well as large changes 9. The complementary primer design results in the mutated plasmid containing staggered nicks, and thus the newly synthesized DNA cannot be used as a template for subsequent amplification 4. The complementary primer pairs favor “primer-dimer” formation by partial annealing of a primer with the second primer in the reaction, instead of primer annealing to the template with mismatches, which causes low PCR amplification efficiency, and may lead to the formation of tandem primer repeats in resulting PCR products and hence a reduction in fidelity 7, 8. The fact that the primers are completely complementary, and hence favor self-annealing limits the PCR product yield and gives rise to false positives 6. coli cells for nick repair.ĭespite its widespread use, the QuikChange system has limitations. The resulting nicked DNA is transformed into competent E. The parental DNA template is eliminated by treating with DpnI, which destroys the methylated template DNA 5. It requires a high-fidelity DNA polymerase that minimizes unwanted mutations, such as KOD hot start DNA polymerase, Pfu DNA polymerase, or Phusion ® high-fidelity DNA polymerase, to amplify the whole plasmid with complementary primer pairs, carrying the desired mutation in the form of mismatches to the original plasmid. Among them, Stratagene’s QuikChange site-directed mutagenesis kit is extremely useful and simple, and probably one of the most favored 4. In the past decade, a number of strategies and commercial kits have been developed for introducing mutational changes in plasmid DNA, such as base substitutions and base additions or deletions. 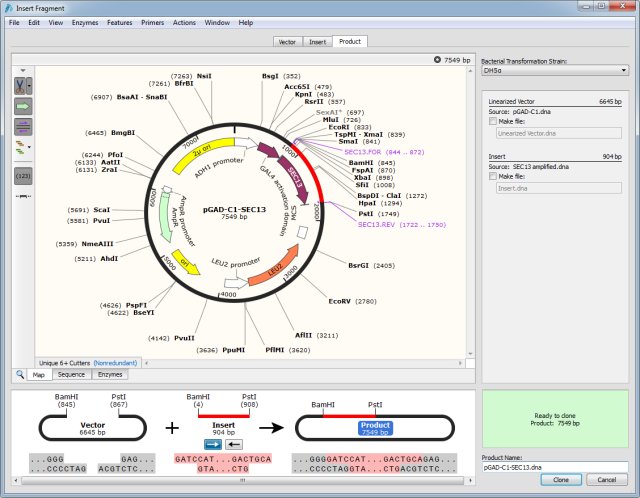
Polymerase chain reaction (PCR)-based site-directed mutagenesis is an invaluable technique for altering genes and hence the structure and activity of individual proteins in a systematic way, opening up opportunities for investigating the structure-function relationships of protein, enzyme specificity and selectivity, or protein engineering 1, 2, 3. LFEAP mutagenesis is a versatile method that offers significant advantages for introducing large and multiple changes in plasmid DNA. With this strategy, multiple simultaneous changes (up to 15) and mutations in large plasmids (up to 50 kb) were achieved with high efficiency and fidelity. These products with compatible sticky ends were efficiently assembled into a circular, mutagenized plasmid. The resulting single strands were then annealed to produce double-stranded DNA with free 5′ single-stranded DNA tails. The first inverse PCR on the target plasmid yielded linearized DNA fragments with mutagenic ends, and a second single primer PCR resulted in complementary single-stranded DNA fragments with the addition of overhangs at the 5′ end of each strand. Here, we propose a new method, named LFEAP mutagenesis ( Ligation of Fragment Ends After PCR) for creating various mutations in plasmid by leveraging three existing concepts: inverse PCR, single primer PCR, and sticky-end assembly. While the QuikChange site-directed mutagenesis method and its later modifications are extremely useful and simple, they suffer from several drawbacks. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
